Find non-linear formulas that fits your input data. You can systematically explore and memoize the possible formulas and its cross-validation performance, in an incremental fashon. Three main interoperable search functions are available:
random.search performs a random exploration,genetic.search employs a genetic optimization
algorithmcomb.search allows the user to provide a data.frame of
formulas to be testedAfter installation, see tutorials by typing
browseVignettes('symbolicr')in your R console.
SymbolicR was featured in the following scientific publication as a data-driven approach to refine and extend a mechanistic model of Fc-fusion protein pharmacokinetics:
Tomasoni et al. Predicting Aberrant Fc-fusion Protein Pharmacokinetics from in Silico Structural Properties and Physiologically Based Pharmacokinetic (PBPK) Modeling, The AAPS Journal, March 2026 link
You can install the development version of symbolicr like so:
devtools::install_github('cosbi-research/symbolicr', build_vignettes = TRUE)This is a minimum viable example on how to use genetic search to find the non-linear relationship:
library(symbolicr)
set.seed(1)
# set-up a toy example dataset
x1<-runif(100, min=2, max=67)
x2<-runif(100, min=0.01, max=0.1)
X <- data.frame(x1=x1, x2=x2)
# set up a "true" non-linear relationship
# with some noise
y <- log10(x1^2*x2) + rnorm(100, 0, 0.001)
results <- genetic.search(
X, y,
n.squares=2,
max.formula.len = 1,
N=2,
K=10,
best.vars.l = list(
c('log.x1')
),
transformations = list(
"log"=function(rdf, x, stats){ log(x) },
"inv"=function(rdf, x, stats){ 1/x }
),
keepBest=T,
cv.norm=F
)
#> ## Total number of single terms: 27
#> Iteration: 1 Mean/Max fitness:-9.80e+07 / 1.66e+00 Best: log.x1
#> Iteration: 2 Mean/Max fitness:-9.80e+07 / 1.66e+00 Best: log.x1
#> Iteration: 3 Mean/Max fitness:-9.80e+07 / 1.66e+00 Best: log.x1
#> Iteration: 4 Mean/Max fitness:-9.60e+07 / 1.66e+00 Best: log.x1
#> Iteration: 5 Mean/Max fitness:-9.40e+07 / 1.66e+00 Best: log.x1
#> Iteration: 6 Mean/Max fitness:-9.40e+07 / 1.66e+00 Best: log.x1
#> Iteration: 7 Mean/Max fitness:-9.40e+07 / 1.66e+00 Best: log.x1
#> Iteration: 8 Mean/Max fitness:-9.40e+07 / 1.66e+00 Best: log.x1
#> Iteration: 9 Mean/Max fitness:-9.60e+07 / 1.66e+00 Best: log.x1
#> Iteration: 10 Mean/Max fitness:-9.60e+07 / 1.66e+00 Best: log.x1
#> Iteration: 11 Mean/Max fitness:-9.60e+07 / 1.66e+00 Best: log.x1
#> Iteration: 12 Mean/Max fitness:-9.60e+07 / 1.66e+00 Best: log.x1
#> Iteration: 13 Mean/Max fitness:-9.40e+07 / 1.66e+00 Best: log.x1
#> Iteration: 14 Mean/Max fitness:-9.40e+07 / 1.66e+00 Best: log.x1
#> Iteration: 15 Mean/Max fitness:-9.60e+07 / 1.66e+00 Best: log.x1
#> Iteration: 16 Mean/Max fitness:-9.40e+07 / 1.66e+00 Best: log.x1
#> Iteration: 17 Mean/Max fitness:-9.60e+07 / 1.66e+00 Best: log.x1
#> Iteration: 18 Mean/Max fitness:-9.40e+07 / 1.66e+00 Best: log.x1
#> Iteration: 19 Mean/Max fitness:-9.00e+07 / 1.69e+00 Best: log.x1
#> Iteration: 20 Mean/Max fitness:-9.60e+07 / 1.69e+00 Best: log.x1
#> Iteration: 21 Mean/Max fitness:-9.00e+07 / 1.69e+00 Best: log.x1
#> Iteration: 22 Mean/Max fitness:-9.40e+07 / 1.69e+00 Best: log.x1
#> Iteration: 23 Mean/Max fitness:-9.40e+07 / 1.69e+00 Best: log.mul.x1.x1
#> Iteration: 24 Mean/Max fitness:-9.60e+07 / 1.69e+00 Best: log.mul.x1.x1
#> Iteration: 25 Mean/Max fitness:-9.60e+07 / 1.69e+00 Best: log.mul.x1.x1
#> Iteration: 26 Mean/Max fitness:-9.40e+07 / 1.71e+00 Best: log.x1
#> Iteration: 27 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 28 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 29 Mean/Max fitness:-9.40e+07 / 1.71e+00 Best: log.x1
#> Iteration: 30 Mean/Max fitness:-9.40e+07 / 1.71e+00 Best: log.x1
#> Iteration: 31 Mean/Max fitness:-9.40e+07 / 1.71e+00 Best: log.x1
#> Iteration: 32 Mean/Max fitness:-9.40e+07 / 1.71e+00 Best: log.x1
#> Iteration: 33 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 34 Mean/Max fitness:-9.40e+07 / 1.71e+00 Best: log.x1
#> Iteration: 35 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 36 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 37 Mean/Max fitness:-9.40e+07 / 1.71e+00 Best: log.x1
#> Iteration: 38 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 39 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 40 Mean/Max fitness:-9.40e+07 / 1.71e+00 Best: log.x1
#> Iteration: 41 Mean/Max fitness:-9.20e+07 / 1.71e+00 Best: log.x1
#> Iteration: 42 Mean/Max fitness:-9.40e+07 / 1.71e+00 Best: log.x1
#> Iteration: 43 Mean/Max fitness:-9.20e+07 / 1.71e+00 Best: log.x1
#> Iteration: 44 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 45 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 46 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 47 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 48 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 49 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 50 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 51 Mean/Max fitness:-9.20e+07 / 1.71e+00 Best: log.x1
#> Iteration: 52 Mean/Max fitness:-9.20e+07 / 1.71e+00 Best: log.x1
#> Iteration: 53 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 54 Mean/Max fitness:-9.40e+07 / 1.71e+00 Best: log.x1
#> Iteration: 55 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 56 Mean/Max fitness:-9.20e+07 / 1.71e+00 Best: log.x1
#> Iteration: 57 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 58 Mean/Max fitness:-9.40e+07 / 1.71e+00 Best: log.x1
#> Iteration: 59 Mean/Max fitness:-9.20e+07 / 1.71e+00 Best: log.x1
#> Iteration: 60 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 61 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 62 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 63 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 64 Mean/Max fitness:-9.40e+07 / 1.71e+00 Best: log.x1
#> Iteration: 65 Mean/Max fitness:-9.40e+07 / 1.71e+00 Best: log.x1
#> Iteration: 66 Mean/Max fitness:-9.60e+07 / 1.71e+00 Best: log.x1
#> Iteration: 67 Mean/Max fitness:-9.40e+07 / 6.22e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 68 Mean/Max fitness:-9.40e+07 / 6.22e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 69 Mean/Max fitness:-9.40e+07 / 6.22e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 70 Mean/Max fitness:-9.20e+07 / 6.22e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 71 Mean/Max fitness:-9.20e+07 / 6.22e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 72 Mean/Max fitness:-9.20e+07 / 6.22e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 73 Mean/Max fitness:-9.40e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 74 Mean/Max fitness:-9.40e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 75 Mean/Max fitness:-9.60e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 76 Mean/Max fitness:-9.40e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 77 Mean/Max fitness:-9.40e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 78 Mean/Max fitness:-9.60e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 79 Mean/Max fitness:-9.60e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 80 Mean/Max fitness:-9.60e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 81 Mean/Max fitness:-9.60e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 82 Mean/Max fitness:-9.60e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 83 Mean/Max fitness:-9.60e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 84 Mean/Max fitness:-9.60e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 85 Mean/Max fitness:-9.40e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 86 Mean/Max fitness:-9.40e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 87 Mean/Max fitness:-9.60e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 88 Mean/Max fitness:-9.60e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 89 Mean/Max fitness:-9.60e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 90 Mean/Max fitness:-9.40e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 91 Mean/Max fitness:-9.40e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 92 Mean/Max fitness:-9.40e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 93 Mean/Max fitness:-9.60e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 94 Mean/Max fitness:-9.40e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 95 Mean/Max fitness:-9.20e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 96 Mean/Max fitness:-9.20e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 97 Mean/Max fitness:-9.20e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 98 Mean/Max fitness:-9.40e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 99 Mean/Max fitness:-9.00e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2
#> Iteration: 100 Mean/Max fitness:-9.20e+07 / 6.23e+00 Best: log.mul.x1.mul.x1.x2We found the correct non-linear formula starting from an initial guess! We can now get the best formula
results$best
#> [1] "log.mul.x1.mul.x1.x2"And all the formula the genetic algorithm found to be best at each one of the 100 evolution iterations (last 5 shown for brevity)
results$best.iter[seq(length(results$best.iter)-5,length(results$best.iter))]
#> [[1]]
#> [1] "log.mul.x1.mul.x1.x2"
#>
#> [[2]]
#> [1] "log.mul.x1.mul.x1.x2"
#>
#> [[3]]
#> [1] "log.mul.x1.mul.x1.x2"
#>
#> [[4]]
#> [1] "log.mul.x1.mul.x1.x2"
#>
#> [[5]]
#> [1] "log.mul.x1.mul.x1.x2"
#>
#> [[6]]
#> [1] "log.mul.x1.mul.x1.x2"Note that cv.norm=FALSE means data is used as-is. Before
running this example we checked for the non-negativeness of
x1^2*x2. If you would like to normalize data to avoid
scaling issues just use cv.norm=TRUE but in this case, to
avoid computing the log of a negative value, we use this updated
transformation function
# NB: function applied to standardized X values!
# they can be negative
log.std <- function(x, stats){ log(0.1 + abs(stats$min) + x) }The stats object contains - min: the minimum of the
column values - absmin: the minimum of the absolute values of the
columns - absmax: the maximum of the absolute values of the columns -
projzero: -mean/sd of the columns, that is the position of the zero in
the original, non-normalized space.
Type ?dataset.min.maxs in your R console for further
informations.
This is a minimum viable example on how to use random search to test multiple non-linear relationships, and get a summary of the performances in a data.frame:
random.results <- random.search(
X, y,
n.squares=2,
formula.len = 1,
N=2,
K=10,
transformations = list(
"log"=function(rdf, x, stats){ log(x) },
"inv"=function(rdf, x, stats){ 1/x }
),
cv.norm=F
)
#> Regression on mul.x2.mul.x2.x2
#> Regression on log.mul.x1.x2
#> Regression on inv.mul.x1.mul.x2.x2
#> Regression on mul.x1.x1
#> Regression on mul.x1.x2
#> Regression on x2
#> Regression on x1
#> Regression on log.x2
#> Regression on mul.x1.mul.x1.x1
#> Regression on log.mul.x2.x2
#> Regression on inv.mul.x2.x2
#> Regression on inv.x2
#> Regression on inv.mul.x1.mul.x1.x2
#> Regression on log.mul.x1.mul.x2.x2
#> Regression on inv.mul.x2.mul.x2.x2
#> Regression on inv.mul.x1.mul.x1.x1
#> Regression on inv.x1
#> Regression on log.mul.x1.mul.x1.x1
#> Regression on mul.x2.x2
#> Regression on log.mul.x1.mul.x1.x2
#> Regression on inv.mul.x1.x2
#> Regression on log.mul.x2.mul.x2.x2
#> Regression on log.mul.x1.x1
#> Regression on inv.mul.x1.x1
#> Regression on mul.x1.mul.x1.x2
#> Regression on log.x1
#> Regression on mul.x1.mul.x2.x2You can then inspect results in the resulting data.frame:
random.results[order(random.results$base.r.squared, decreasing = T), ][seq(5), ]
#> base.pe base.cor base.r.squared base.max.pe base.iqr.pe
#> 20 0.0004285218 0.9999988 0.9999976 0.004748987 0.0005285826
#> 2 0.0725826920 0.9536868 0.9095013 1.433554884 0.1219353333
#> 26 0.1038707364 0.9276918 0.8606065 1.093821498 0.0978448963
#> 23 0.1005819365 0.9272320 0.8597596 1.111784453 0.0969222114
#> 18 0.1039579306 0.9261947 0.8578210 1.085202757 0.0888076193
#> base.max.cooksd base.max.cooksd.name vars n.squares
#> 20 0.4515147 47 log.mul.x1.mul.x1.x2 2
#> 2 0.2559093 27 log.mul.x1.x2 2
#> 26 0.1554620 47 log.x1 2
#> 23 0.1542245 47 log.mul.x1.x1 2
#> 18 0.1567286 47 log.mul.x1.mul.x1.x1 2
#> formula.len
#> 20 1
#> 2 1
#> 26 1
#> 23 1
#> 18 1Search over all non-duplicated combinations of terms taken from an user-supplied list of formulas.
# test the three formula x1, x2 and log(x2)
comb.search(
X, y,
combinations=data.frame(t1=c('x1','x2','log.x2')),
N=2,
K=10,
transformations = list(
"log"=function(rdf, x, stats){ log(x) },
"exp"=function(rdf, x, stats){ exp(x) }
),
cv.norm=F
)
#> Regression on x1
#> Regression on x2
#> Regression on log.x2
#> base.pe base.cor base.r.squared base.max.pe base.iqr.pe base.max.cooksd
#> 1 0.1302693 0.8630977 0.74487976 3.080926 0.1708999 0.3215164
#> 2 0.2173972 0.3203810 0.09942148 7.524692 0.1991448 0.1019341
#> 3 0.2215673 0.2952262 0.08528910 7.786758 0.1758801 0.1168211
#> base.max.cooksd.name vars n.squares formula.len
#> 1 47 x1 0 1
#> 2 47 x2 0 1
#> 3 47 log.x2 0 1This code highlights the most-important variables among a list of formulas applied to a given dataset.
max.formula.len=1
transformations=list(
"log"=function(rdf, x, stats){ log(x) },
"log_x1_p"=function(rdf, x, stats){ log(rdf$x1 + x) },
"inv"=function(rdf, x, stats){ 1/x }
)
random.results <- random.search(
X, y,
n.squares=2,
formula.len = max.formula.len,
N=2,
K=10,
transformations = transformations,
cv.norm=F
)
#> Regression on mul.x1.x1
#> Regression on log.mul.x1.x1
#> Regression on mul.x2.mul.x2.x2
#> Regression on mul.x2.x2
#> Regression on inv.mul.x1.mul.x2.x2
#> Regression on log_x1_p.x1
#> Regression on inv.mul.x2.mul.x2.x2
#> Regression on inv.mul.x1.x1
#> Regression on mul.x1.mul.x1.x1
#> Regression on log_x1_p.mul.x1.mul.x2.x2
#> Regression on inv.mul.x2.x2
#> Regression on inv.x1
#> Regression on inv.x2
#> Regression on mul.x1.mul.x2.x2
#> Regression on x1
#> Regression on mul.x1.mul.x1.x2
#> Regression on x2
#> Regression on mul.x1.x2
#> Regression on log_x1_p.mul.x1.x2
#> Regression on log.mul.x1.mul.x1.x2
#> Regression on log_x1_p.mul.x1.x1
#> Regression on inv.mul.x1.mul.x1.x2
#> Regression on log.x1
#> Regression on inv.mul.x1.x2
#> Regression on log.x2
#> Regression on log_x1_p.mul.x2.mul.x2.x2
#> Regression on inv.mul.x1.mul.x1.x1
#> Regression on log_x1_p.mul.x1.mul.x1.x2
#> Regression on log_x1_p.x2
#> Regression on log_x1_p.mul.x2.x2
#> Regression on log.mul.x1.x2
#> Regression on log.mul.x2.x2
#> Regression on log.mul.x1.mul.x2.x2
#> Regression on log.mul.x1.mul.x1.x1
#> Regression on log_x1_p.mul.x1.mul.x1.x1
#> Regression on log.mul.x2.mul.x2.x2
# compute a unique objective function
random.results$obj <- apply(random.results, MARGIN=1, FUN=function(row) pe.r.squared.formula.len.fitness(as.data.frame(t(row)), max.formula.len))
# sort by top-N functions (according to obj)
ordered.res <- random.results[order(random.results$obj,decreasing=T),]
# max fitness on all computed formulas
best.obj <- ordered.res[1,'obj']
# analyze top-10 formulas
# select formulas according to criterion above
eligible.res <- ordered.res[seq(10), ]
direction = 'max' # obj should be maximized
sensitivity <- analyze.variables(
X, y, eligible.res, fitness.column='obj',
# a list of available term transformations
transformations=transformations,
# a list of rules to remove a variable from a term
# ex.
# orig transformation -> base transformation, removed term
# "log_empty_well_p"=c("log10", "empty_well")
transformations_replacement_map=list(
"log_x1_p"=c("log", "x1")
),
custom.abs.mins=list(),
K=10,
N=2,
direction=direction,
max.formula.len=max.formula.len,
fitness.fun=pe.r.squared.formula.len.fitness,
cv.norm=F
)
#> Analyzing log.mul.x1.mul.x1.x2
#> Regression on log.mul.x1.x2
#> Regression on log.mul.x1.x2
#> Regression on log.mul.x1.x1
#> Analyzing log_x1_p.mul.x1.mul.x1.x2
#> Regression on log.mul.x1.mul.x1.x2
#> Regression on log_x1_p.mul.x1.x2
#> Regression on log_x1_p.mul.x1.x2
#> Regression on log_x1_p.mul.x1.x1
#> Analyzing log.mul.x1.x2
#> Regression on log.x2
#> Regression on log.x1
#> Analyzing log_x1_p.mul.x1.x2
#> Regression on log.mul.x1.x2
#> Regression on log_x1_p.x2
#> Regression on log_x1_p.x1
#> Analyzing log_x1_p.x2
#> Regression on log.x2
#> Analyzing log_x1_p.mul.x1.mul.x2.x2
#> Regression on log.mul.x1.mul.x2.x2
#> Regression on log_x1_p.mul.x2.x2
#> Regression on log_x1_p.mul.x1.x2
#> Regression on log_x1_p.mul.x1.x2
#> Warning in rbind(deparse.level, ...): number of columns of result, 3, is not a
#> multiple of vector length 2 of arg 1
#> Analyzing log_x1_p.x1
#> Regression on log.x1
#> Analyzing log.mul.x1.mul.x1.x1
#> Regression on log.mul.x1.x1
#> Regression on log.mul.x1.x1
#> Regression on log.mul.x1.x1
#> Analyzing log_x1_p.mul.x1.mul.x1.x1
#> Regression on log.mul.x1.mul.x1.x1
#> Regression on log_x1_p.mul.x1.x1
#> Regression on log_x1_p.mul.x1.x1
#> Regression on log_x1_p.mul.x1.x1
#> Warning in rbind(deparse.level, ...): number of columns of result, 3, is not a
#> multiple of vector length 4 of arg 1
#> Analyzing log_x1_p.mul.x2.mul.x2.x2
#> Regression on log.mul.x2.mul.x2.x2
#> Regression on log_x1_p.mul.x2.x2
#> Regression on log_x1_p.mul.x2.x2
#> Regression on log_x1_p.mul.x2.x2
# plottable data.frame of quantile losses per-variable
variable.importance.df <- sensitivity[['var.imp']]
n.formulas <- 10
#### PLOT ANALYSIS ####
library(ggplot2)
#> Warning: package 'ggplot2' was built under R version 4.5.2
library(RColorBrewer)
library(colorspace)
#> Warning: package 'colorspace' was built under R version 4.5.2
colours <- brewer.pal(11, "Paired")
cols_d4 <- darken(colours, 0.4)
#png(file.path('sensitivity', paste("sensitivity.",(percent.limit*100),"percent",type,".png", sep = "")), width = 640, height = 490)
p <- ggplot(variable.importance.df, aes(x=mean.occurrences, y=mean.loss.p, colour=variable)) +
theme(
text = element_text(size = 15)
) +
labs(x = paste0("Mean number of occurrences for best ",n.formulas," formulas"), y = "relative loss %") +
# geom_crossbar(aes(ymin = lower.loss.p, ymax = higher.loss.p), width = 1.5) +
geom_pointrange(aes(ymin = lower.loss.p, ymax = higher.loss.p)) +
scale_y_continuous(trans = 'log10', limits=c(1,110), n.breaks=8,
labels =c("1%","3%","5%", "10%", "20%", "30%", "50%", "100%"),
breaks =c(1, 3, 5, 10, 20, 30, 50, 100))+
scale_x_continuous(limits=c(0,3.05), n.breaks=6,
labels =c("1/5", "1/2", "3/4", "1", "2", "3"),
breaks =c(1/5.0, 1/2.0, 3/4.0, 1, 2, 3))+
scale_color_manual(values = cols_d4)
print(p)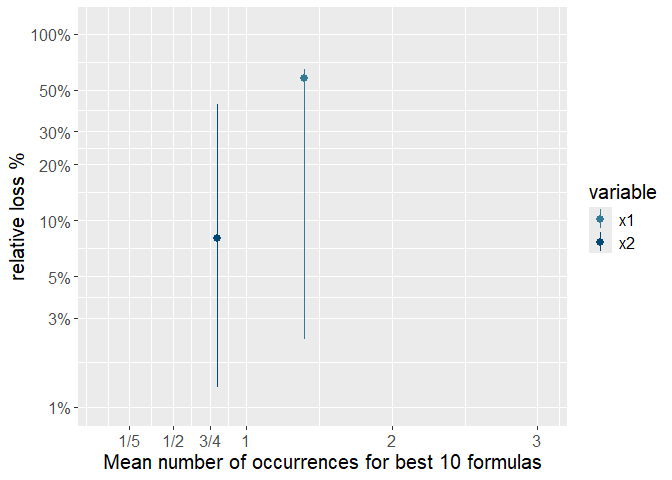
This plot is showing clearly that variable x1 is much more important than variable x2 as it both